Plot variable importance scores for the predictors in a model.
Usage
vip(object, ...)
# Default S3 method
vip(
object,
num_features = 10L,
geom = c("col", "point", "boxplot", "violin"),
mapping = NULL,
aesthetics = list(),
horizontal = TRUE,
all_permutations = FALSE,
jitter = FALSE,
include_type = FALSE,
...
)
# S3 method for class 'model_fit'
vip(object, ...)
# S3 method for class 'workflow'
vip(object, ...)
# S3 method for class 'WrappedModel'
vip(object, ...)
# S3 method for class 'Learner'
vip(object, ...)Arguments
- object
A fitted model (e.g., of class randomForest object) or a vi object.
- ...
Additional optional arguments to be passed on to vi.
- num_features
Integer specifying the number of variable importance scores to plot. Default is
10.- geom
Character string specifying which type of plot to construct. The currently available options are described below.
geom = "col"uses geom_col to construct a bar chart of the variable importance scores.geom = "point"uses geom_point to construct a Cleveland dot plot of the variable importance scores.geom = "boxplot"uses geom_boxplot to construct a boxplot plot of the variable importance scores. This option can only for the permutation-based importance method withnsim > 1andkeep = TRUE; see vi_permute for details.geom = "violin"uses geom_violin to construct a violin plot of the variable importance scores. This option can only for the permutation-based importance method withnsim > 1andkeep = TRUE; see vi_permute for details.
- mapping
Set of aesthetic mappings created by aes-related functions and/or tidy eval helpers. See example usage below.
- aesthetics
List specifying additional arguments passed on to layer. These are often aesthetics, used to set an aesthetic to a fixed value, like
colour = "red"orsize = 3. See example usage below.- horizontal
Logical indicating whether or not to plot the importance scores on the x-axis (
TRUE). Default isTRUE.- all_permutations
Logical indicating whether or not to plot all permutation scores along with the average. Default is
FALSE. (Only used for permutation scores whennsim > 1.)- jitter
Logical indicating whether or not to jitter the raw permutation scores. Default is
FALSE. (Only used whenall_permutations = TRUE.)- include_type
Logical indicating whether or not to include the type of variable importance computed in the axis label. Default is
FALSE.
Examples
#
# A projection pursuit regression example using permutation-based importance
#
# Load the sample data
data(mtcars)
# Fit a projection pursuit regression model
model <- ppr(mpg ~ ., data = mtcars, nterms = 1)
# Construct variable importance plot (permutation importance, in this case)
set.seed(825) # for reproducibility
pfun <- function(object, newdata) predict(object, newdata = newdata)
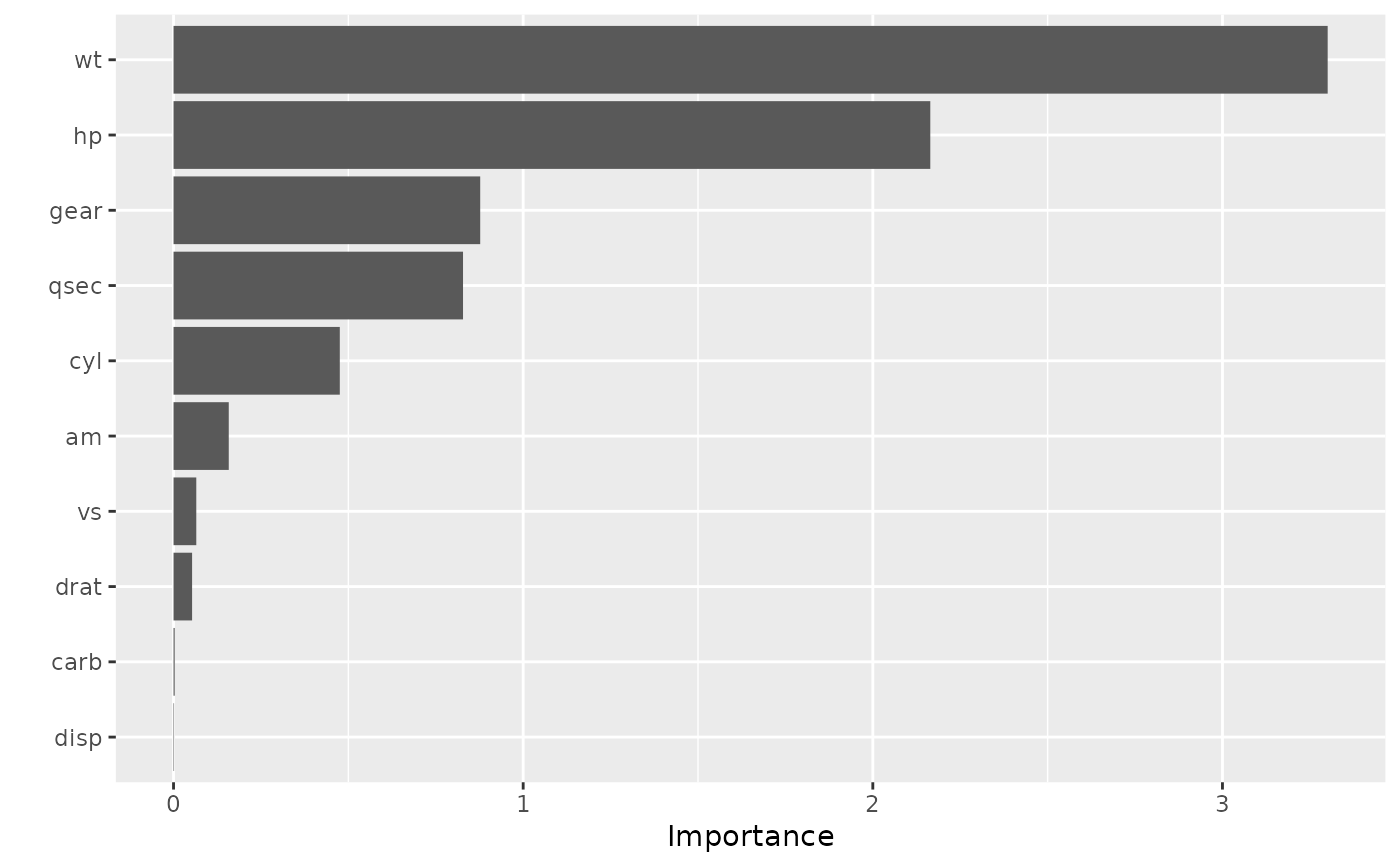
vip(model, method = "permute", train = mtcars, target = "mpg", nsim = 10,
metric = "rmse", pred_wrapper = pfun)
 # Better yet, store the variable importance scores and then plot
set.seed(825) # for reproducibility
vis <- vi(model, method = "permute", train = mtcars, target = "mpg",
nsim = 10, metric = "rmse", pred_wrapper = pfun)
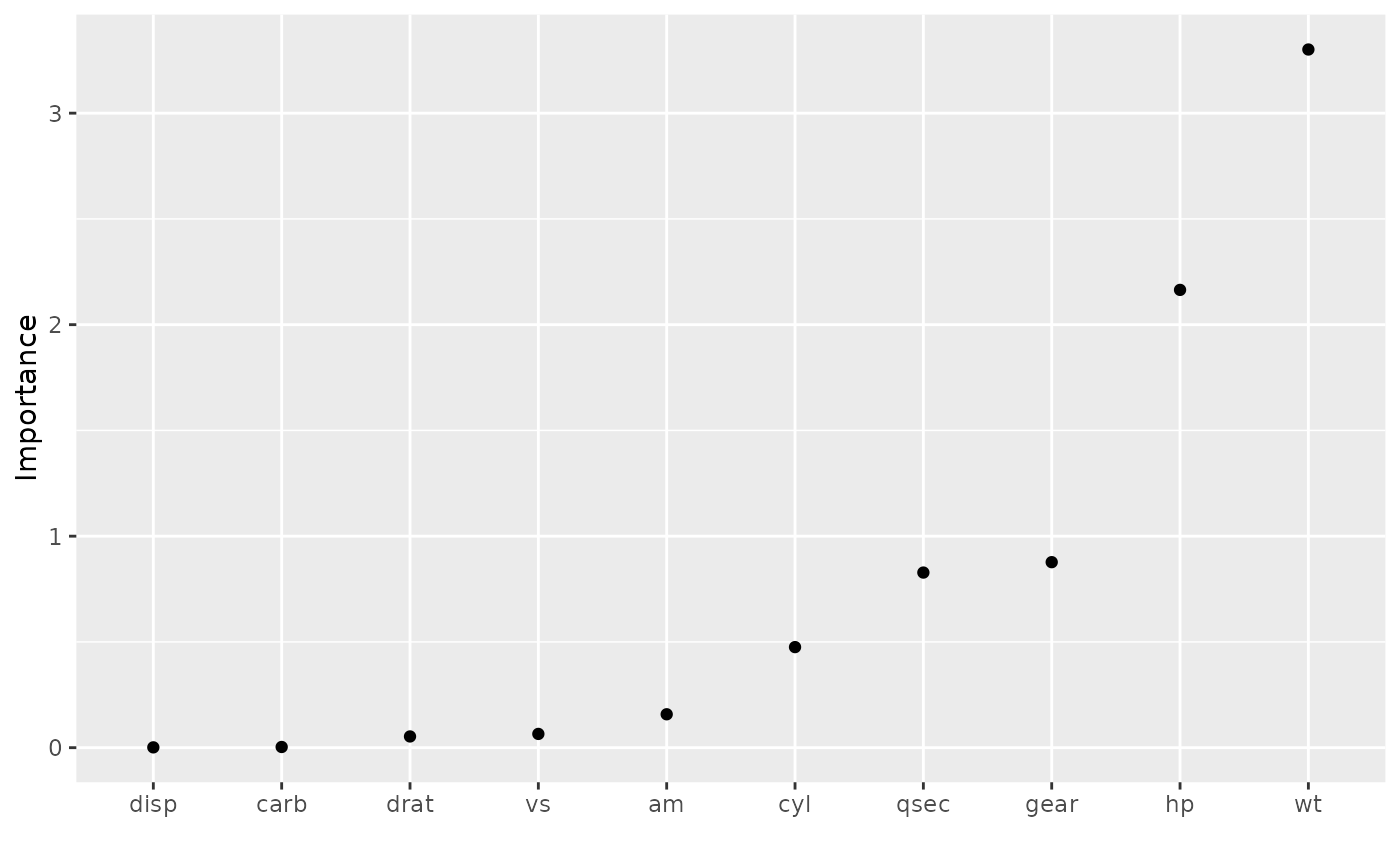
vip(vis, geom = "point", horiz = FALSE)
# Better yet, store the variable importance scores and then plot
set.seed(825) # for reproducibility
vis <- vi(model, method = "permute", train = mtcars, target = "mpg",
nsim = 10, metric = "rmse", pred_wrapper = pfun)
vip(vis, geom = "point", horiz = FALSE)
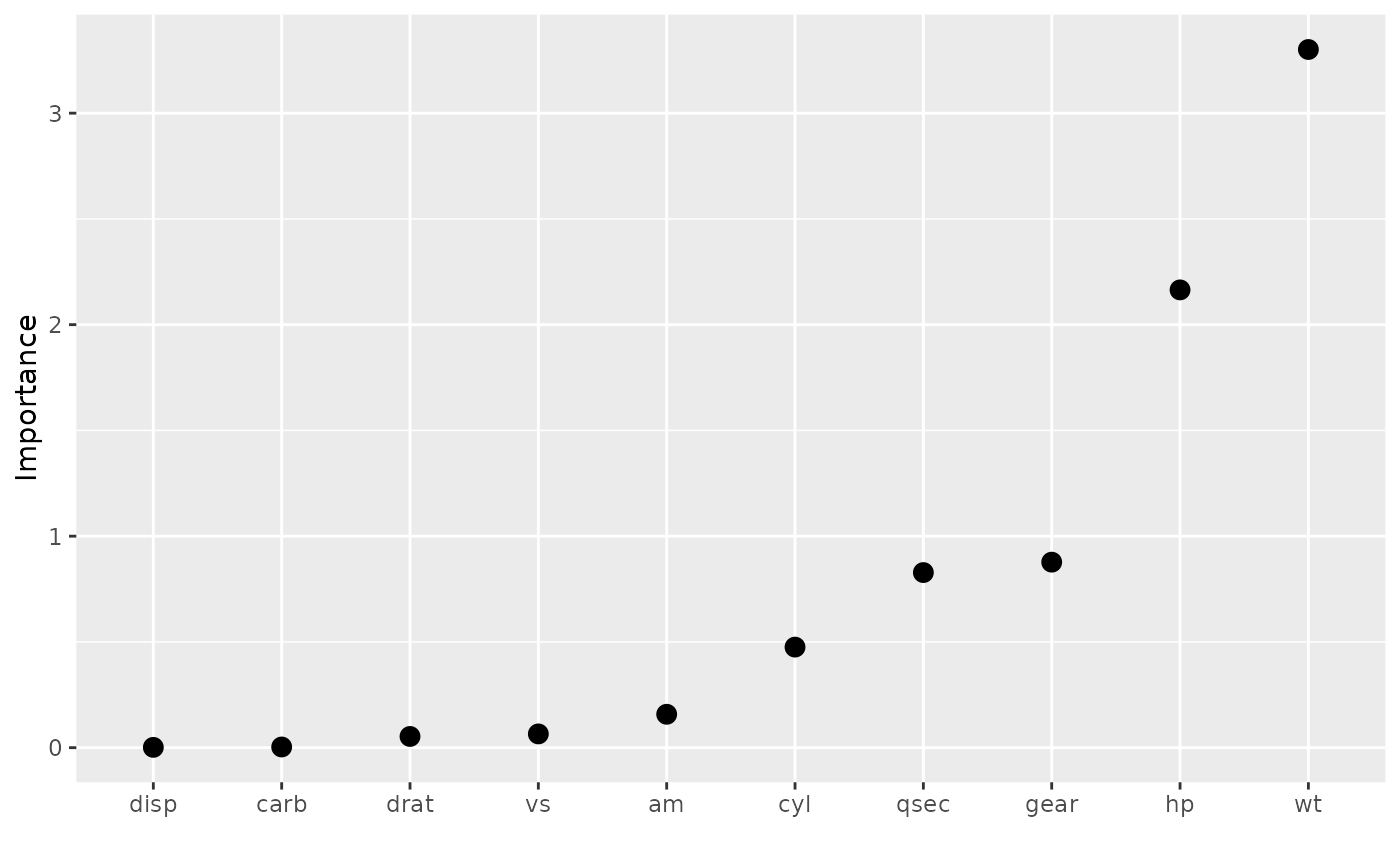
 vip(vis, geom = "point", horiz = FALSE, aesthetics = list(size = 3))
vip(vis, geom = "point", horiz = FALSE, aesthetics = list(size = 3))
 # Plot unaggregated permutation scores (boxplot colored by feature)
library(ggplot2) # for `aes()`-related functions and tidy eval helpers
vip(vis, geom = "boxplot", all_permutations = TRUE, jitter = TRUE,
#mapping = aes_string(fill = "Variable"), # for ggplot2 (< 3.0.0)
mapping = aes(fill = .data[["Variable"]]), # for ggplot2 (>= 3.0.0)
aesthetics = list(color = "grey35", size = 0.8))
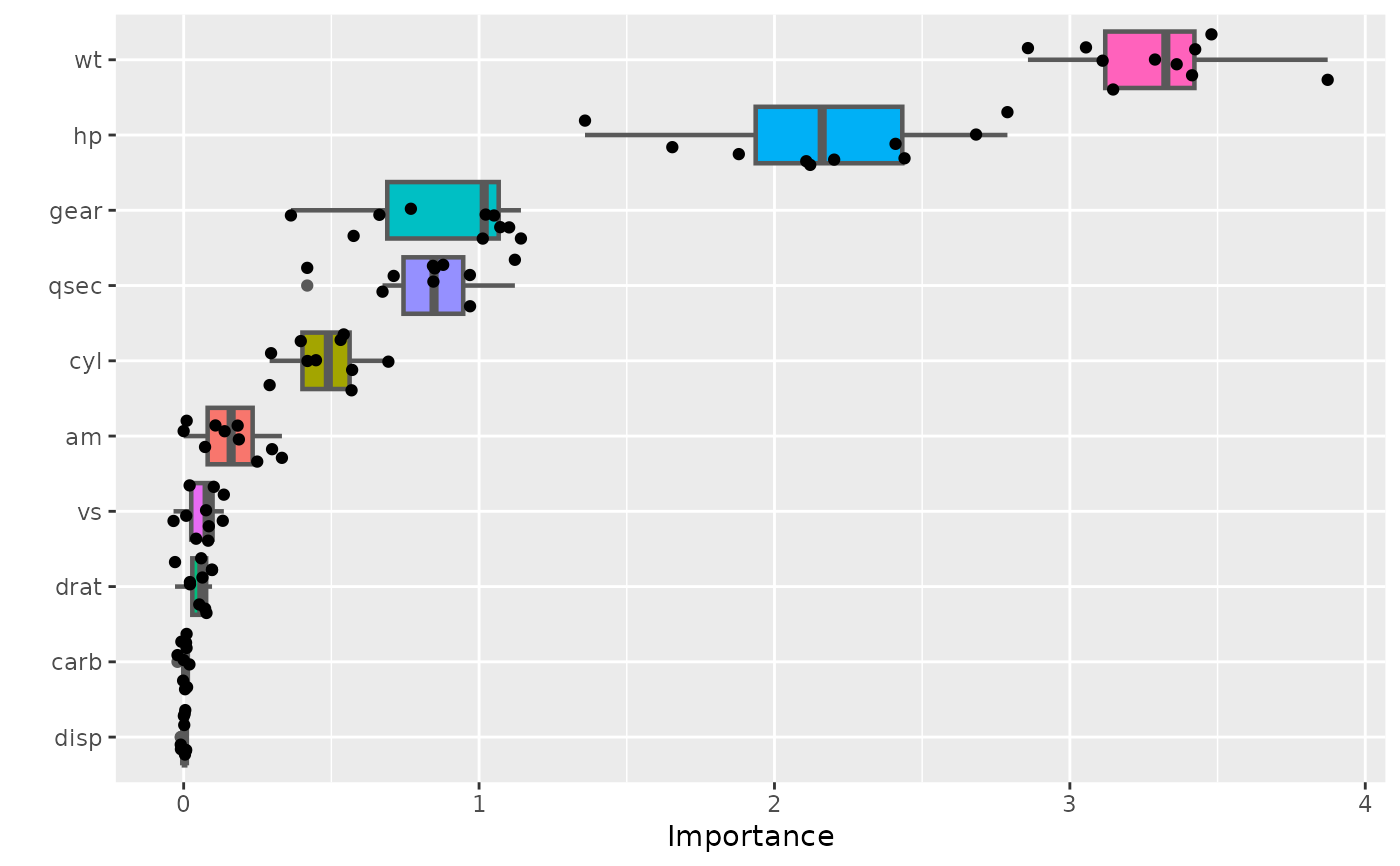
# Plot unaggregated permutation scores (boxplot colored by feature)
library(ggplot2) # for `aes()`-related functions and tidy eval helpers
vip(vis, geom = "boxplot", all_permutations = TRUE, jitter = TRUE,
#mapping = aes_string(fill = "Variable"), # for ggplot2 (< 3.0.0)
mapping = aes(fill = .data[["Variable"]]), # for ggplot2 (>= 3.0.0)
aesthetics = list(color = "grey35", size = 0.8))
 #
# A binary classification example
#
if (FALSE) { # \dontrun{
library(rpart) # for classification and regression trees
# Load Wisconsin breast cancer data; see ?mlbench::BreastCancer for details
data(BreastCancer, package = "mlbench")
bc <- subset(BreastCancer, select = -Id) # for brevity
# Fit a standard classification tree
set.seed(1032) # for reproducibility
tree <- rpart(Class ~ ., data = bc, cp = 0)
# Prune using 1-SE rule (e.g., use `plotcp(tree)` for guidance)
cp <- tree$cptable
cp <- cp[cp[, "nsplit"] == 2L, "CP"]
tree2 <- prune(tree, cp = cp) # tree with three splits
# Default tree-based VIP
vip(tree2)
# Computing permutation importance requires a prediction wrapper. For
# classification, the return value depends on the chosen metric; see
# `?vip::vi_permute` for details.
pfun <- function(object, newdata) {
# Need vector of predicted class probabilities when using log-loss metric
predict(object, newdata = newdata, type = "prob")[, "malignant"]
}
# Permutation-based importance (note that only the predictors that show up
# in the final tree have non-zero importance)
set.seed(1046) # for reproducibility
vip(tree2, method = "permute", nsim = 10, target = "Class",
metric = "logloss", pred_wrapper = pfun, reference_class = "malignant")
} # }
#
# A binary classification example
#
if (FALSE) { # \dontrun{
library(rpart) # for classification and regression trees
# Load Wisconsin breast cancer data; see ?mlbench::BreastCancer for details
data(BreastCancer, package = "mlbench")
bc <- subset(BreastCancer, select = -Id) # for brevity
# Fit a standard classification tree
set.seed(1032) # for reproducibility
tree <- rpart(Class ~ ., data = bc, cp = 0)
# Prune using 1-SE rule (e.g., use `plotcp(tree)` for guidance)
cp <- tree$cptable
cp <- cp[cp[, "nsplit"] == 2L, "CP"]
tree2 <- prune(tree, cp = cp) # tree with three splits
# Default tree-based VIP
vip(tree2)
# Computing permutation importance requires a prediction wrapper. For
# classification, the return value depends on the chosen metric; see
# `?vip::vi_permute` for details.
pfun <- function(object, newdata) {
# Need vector of predicted class probabilities when using log-loss metric
predict(object, newdata = newdata, type = "prob")[, "malignant"]
}
# Permutation-based importance (note that only the predictors that show up
# in the final tree have non-zero importance)
set.seed(1046) # for reproducibility
vip(tree2, method = "permute", nsim = 10, target = "Class",
metric = "logloss", pred_wrapper = pfun, reference_class = "malignant")
} # }